A Formal Introduction to the Amino Acids
When you Google “amino acids,” one of the first images you’ll see is this colorful diagram, showing the skeletal formulas of twenty molecules grouped by the chemical character of their side chains. Throughout many biochemistry courses, I have found myself memorizing this exact chart.
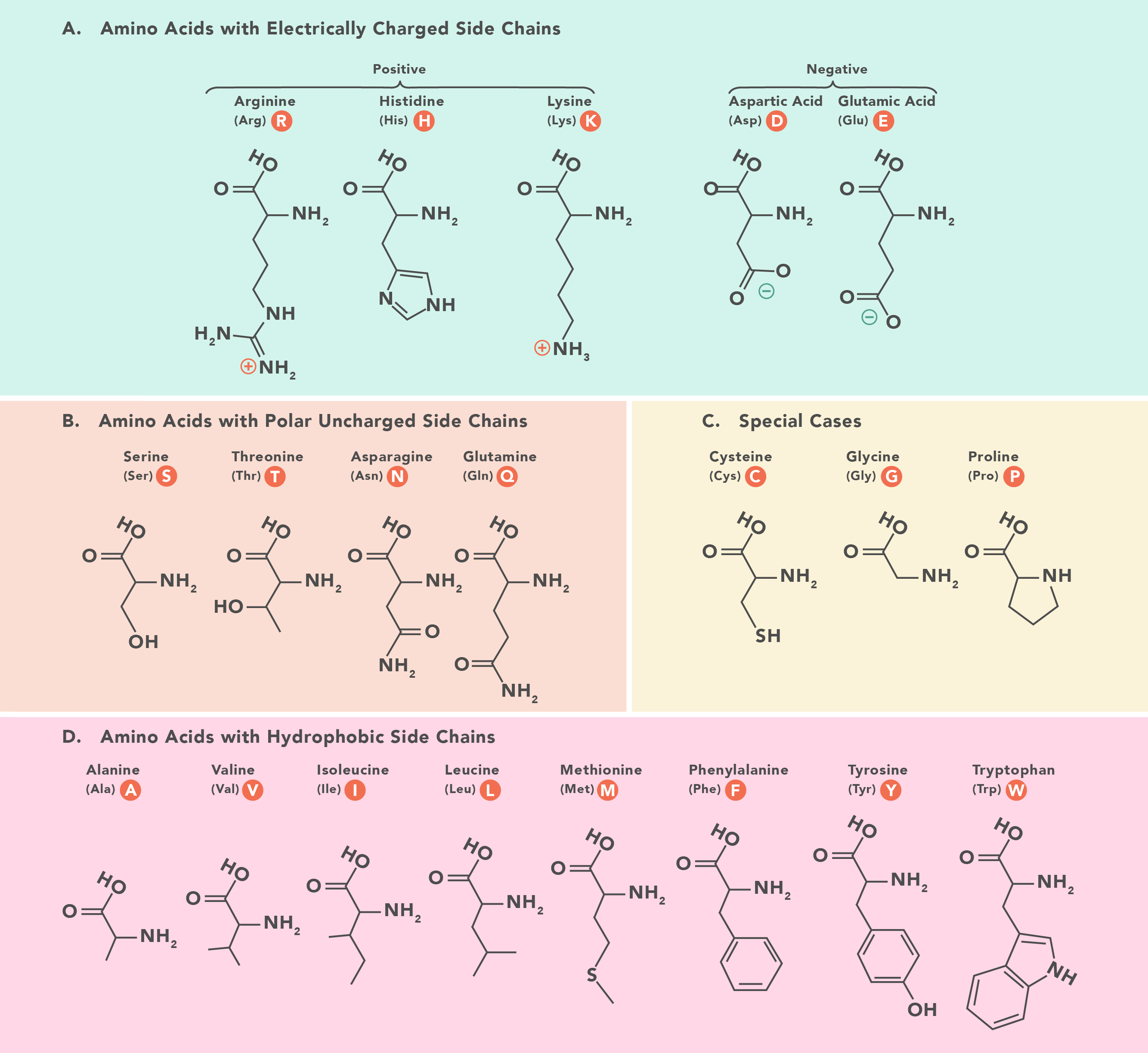
I’ve often glanced these over and moved on to higher-level concepts like enzyme mechanics or structure analysis, but this time I wanted to appreciate the amino acids a bit more and use this as an excuse to try the blog feature of this website.
A brief aside on the amino acid synthetases
There are over 800 naturally occurring amino acids humans have found in nature, but only 20 are directly encoded by the genetic code and incorporated into proteins during translation. These are the proteinogenic amino acids. What distinguishes them from the other 780+ is that each one is selected for by one of the 20 unique aminoacyl-tRNA synthetases (aaRS), each of which loads a specific amino acid onto the corresponding transfer RNAs (tRNAs), specified by their codons.
In my experience, the synthetases are often overlooked in introductory courses, but are arguably among the most important enzymes in molecular biology. They are among the oldest enzymes in cells, predating the divergence of the domains of life. The 20 synthetases fall into two structurally unrelated classes (Class I and Class II), each with 10 members, that likely evolved independently. Despite having completely different protein folds, both classes solve the same problem: recognizing one amino acid out of twenty and attaching it to the right tRNA with an overall translation error rate below 1 in 10,000. Consider that some amino acids are chemically near-identical: valine and isoleucine differ by the placement of a single methyl group, yet the synthetases reliably distinguish them. Several synthetases have a dedicated proofreading domain, a second active site that hydrolyzes incorrectly charged tRNAs before the mischarged tRNA can reach the ribosome. The accuracy of the synthetases is so exceptional that they are distinguished with the term “superspecificity”. Not all synthetases have a dedicated editing domain; those that lack one compensate through highly specific binding and activation of their cognate amino acid. Interestingly, the accuracy of these attachments also depends on stoichiometry: cells must maintain a precise ratio of each synthetase to its cognate tRNAs, because overproduction of a synthetase leads to increased misacylation (the wrong amino acid is attached to the wrong tRNA).
61 sense codons (out of $4^3=64$ total, with 3 stop codons) map to 20 amino acids, mediated by roughly 45 distinct tRNA species and exactly 20 synthetases, one for each amino acid (plus a 21st, selenocysteine, which we’ll get to in the next section).
The $21^*$ Proteinogenic Amino Acids
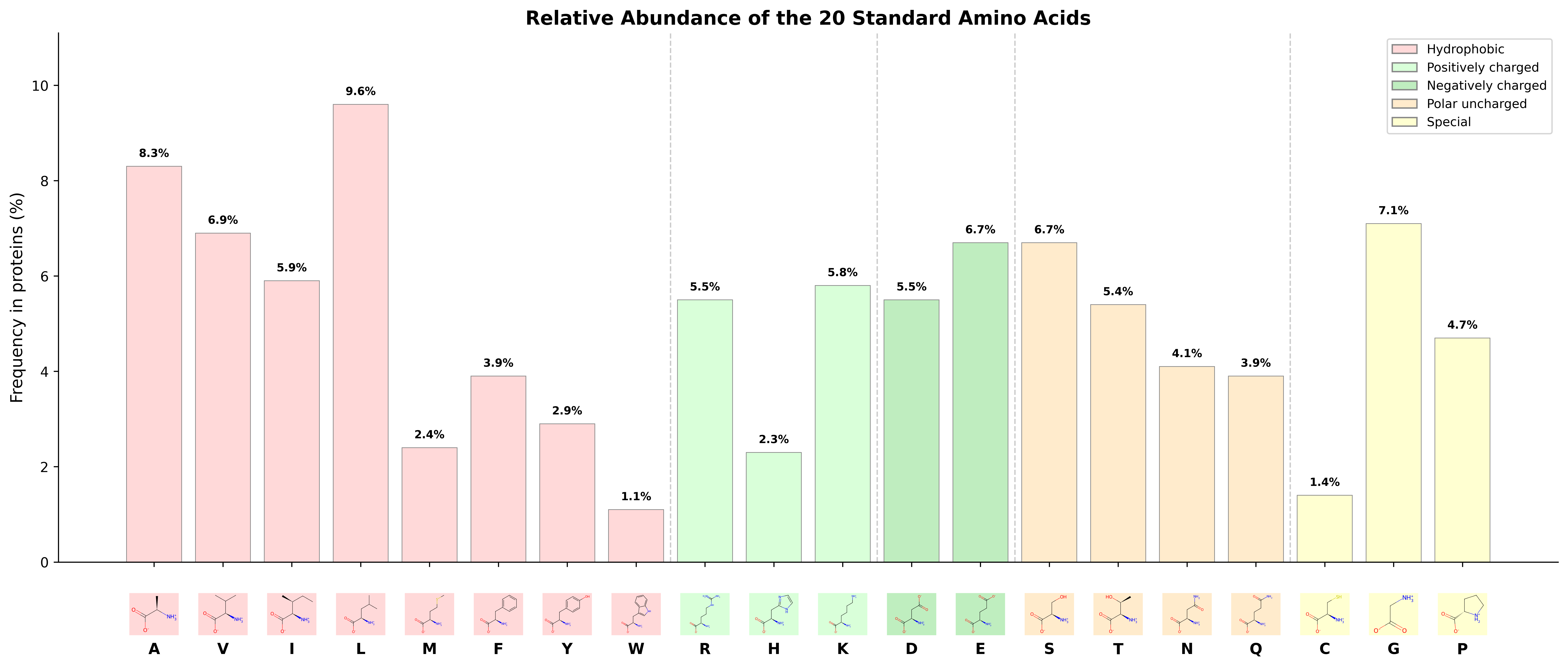
Hydrophobic Side Chains
The largest group contains eight amino acids whose side chains are made mostly or entirely of carbon and hydrogen, making them nonpolar and hydrophobic. In a folded protein, these residues tend to cluster in the interior, away from water, forming the hydrophobic core that drives protein folding.
Alanine

| Charge | Neutral |
| Heavy R atoms | 1 |
| Frequency | ~8.3% |
Valine

| Charge | Neutral |
| Heavy R atoms | 3 |
| Frequency | ~6.9% |
Isoleucine

| Charge | Neutral |
| Heavy R atoms | 4 |
| Frequency | ~5.9% |
Leucine

| Charge | Neutral |
| Heavy R atoms | 4 |
| Frequency | ~9.6% |
Methionine

| Charge | Neutral |
| Heavy R atoms | 4 |
| Frequency | ~2.4% |
Phenylalanine

| Charge | Neutral |
| Heavy R atoms | 7 |
| Frequency | ~3.9% |
Tyrosine

| Charge | Neutral |
| Heavy R atoms | 8 |
| Frequency | ~2.9% |
Tryptophan

| Charge | Neutral |
| Heavy R atoms | 10 |
| Frequency | ~1.1% |
Electrically Charged Side Chains
Five amino acids carry a net charge at physiological pH. Three are positively charged (arginine, histidine, lysine) and two are negatively charged (aspartate, glutamate). These residues are almost always found on protein surfaces, where they interact with water, form salt bridges with oppositely charged residues, and participate in catalysis.
Arginine

| Charge | +1 |
| Heavy R atoms | 7 |
| Frequency | ~5.5% |
Histidine

| Charge | ~0 (pKa ~6.0) |
| Heavy R atoms | 5 |
| Frequency | ~2.3% |
Lysine

| Charge | +1 |
| Heavy R atoms | 5 |
| Frequency | ~5.8% |
Aspartate

| Charge | -1 |
| Heavy R atoms | 3 |
| Frequency | ~5.5% |
Glutamate

| Charge | -1 |
| Heavy R atoms | 4 |
| Frequency | ~6.7% |
Polar Uncharged Side Chains
These four amino acids have side chains that can form hydrogen bonds with water and other polar groups, but carry no net charge at physiological pH. This makes them common on protein surfaces and at active sites, where hydrogen bonding is critical.
Serine

| Charge | Neutral |
| Heavy R atoms | 2 |
| Frequency | ~6.7% |
Threonine

| Charge | Neutral |
| Heavy R atoms | 3 |
| Frequency | ~5.4% |
Asparagine

| Charge | Neutral |
| Heavy R atoms | 4 |
| Frequency | ~4.1% |
Glutamine

| Charge | Neutral |
| Heavy R atoms | 5 |
| Frequency | ~3.9% |
Special Cases
These three amino acids have unusual structural properties that set them apart from the other groups.
Cysteine

| Charge | Neutral |
| Heavy R atoms | 2 |
| Frequency | ~1.4% |
Glycine

| Charge | Neutral |
| Heavy R atoms | 0 |
| Frequency | ~7.1% |
Proline

| Charge | Neutral |
| Heavy R atoms | 3 |
| Frequency | ~4.7% |
The 21st Proteinogenic Amino Acid
Selenocysteine (Sec, U) is a structural analog of cysteine with a selenium atom in place of sulfur. It is found across all three domains of life but is not universal (fungi and higher plants have lost it, for example). What makes it remarkable is how it ends up in proteins. Every other proteinogenic amino acid is specified by one or more sense codons, delivered by a dedicated tRNA, and charged by its own aminoacyl-tRNA synthetase. Selenocysteine has none of these things.
Selenocysteine has neither a codon of its own nor a dedicated aminoacyl-tRNA synthetase. It is encoded by UGA, normally one of three stop codons, and the specialized tRNA^Sec that carries it is first charged with serine by seryl-tRNA synthetase, then converted to selenocysteine directly on the tRNA by selenocysteine synthase. And the ribosome only recodes UGA from “stop” to “selenocysteine” when the mRNA contains a downstream stem-loop called a SECIS element (selenocysteine insertion sequence), which is bound by a specialized elongation factor (SelB in bacteria, EFSec in eukaryotes) that recruits the charged tRNA^Sec to the UGA codon.
Selenocysteine

| Charge | Neutral |
| Heavy R atoms | 2 |
| Frequency | < 0.01% |
Substitution Matrices
Proteins evolve, and when we compare homologous proteins from different species we find that some amino acids substitute for each other constantly while others almost never do. The pattern is not random; it reflects which substitutions the protein can tolerate, which in turn reflects the physicochemical similarity of the amino acids involved.
The standard way to quantify this is a substitution matrix: a 20×20 table where each entry is a score for how likely it is that one amino acid replaces another in evolution. Positive scores mean “more common than chance” (the substitution is tolerated), and negative scores mean “less common than chance” (the substitution is avoided). The most widely used is BLOSUM62, derived in 1992 by Steven and Jorja Henikoff from conserved blocks of homologous protein sequences. The “62” refers to the clustering threshold: before counting substitutions, sequence pairs with ≥62% identity were clustered together so that closely-related sequences wouldn’t dominate the statistics. BLOSUM62 is the default matrix in BLAST and in most protein alignment tools.
BLOSUM scores aren’t arbitrary numbers; they’re log-odds ratios. The score for substituting amino acid $i$ with amino acid $j$ is:
\[S_{ij} = \frac{1}{\lambda} \log_2 \frac{p_{ij}}{q_i q_j}\]where $p_{ij}$ is the observed probability that $i$ and $j$ appear aligned in conserved blocks of homologous proteins, $q_i$ and $q_j$ are the background frequencies of each amino acid in the dataset, and $\lambda$ is a scaling factor chosen to make the final scores convenient integers.
The ratio $p_{ij} / (q_i q_j)$ compares the observed substitution frequency to what you’d expect if the two amino acids paired up purely by chance. A positive $S_{ij}$ means the substitution is more common than chance (the pair co-occurs in conserved positions more often than random pairing would predict), a negative $S_{ij}$ means it’s rarer than chance (evolution avoids it), and a score of zero means observed matches expected. Taking the log turns this ratio into an additive score, so that when you score an alignment of two sequences, you can simply sum the per-position scores to get a total log-odds score for the alignment.
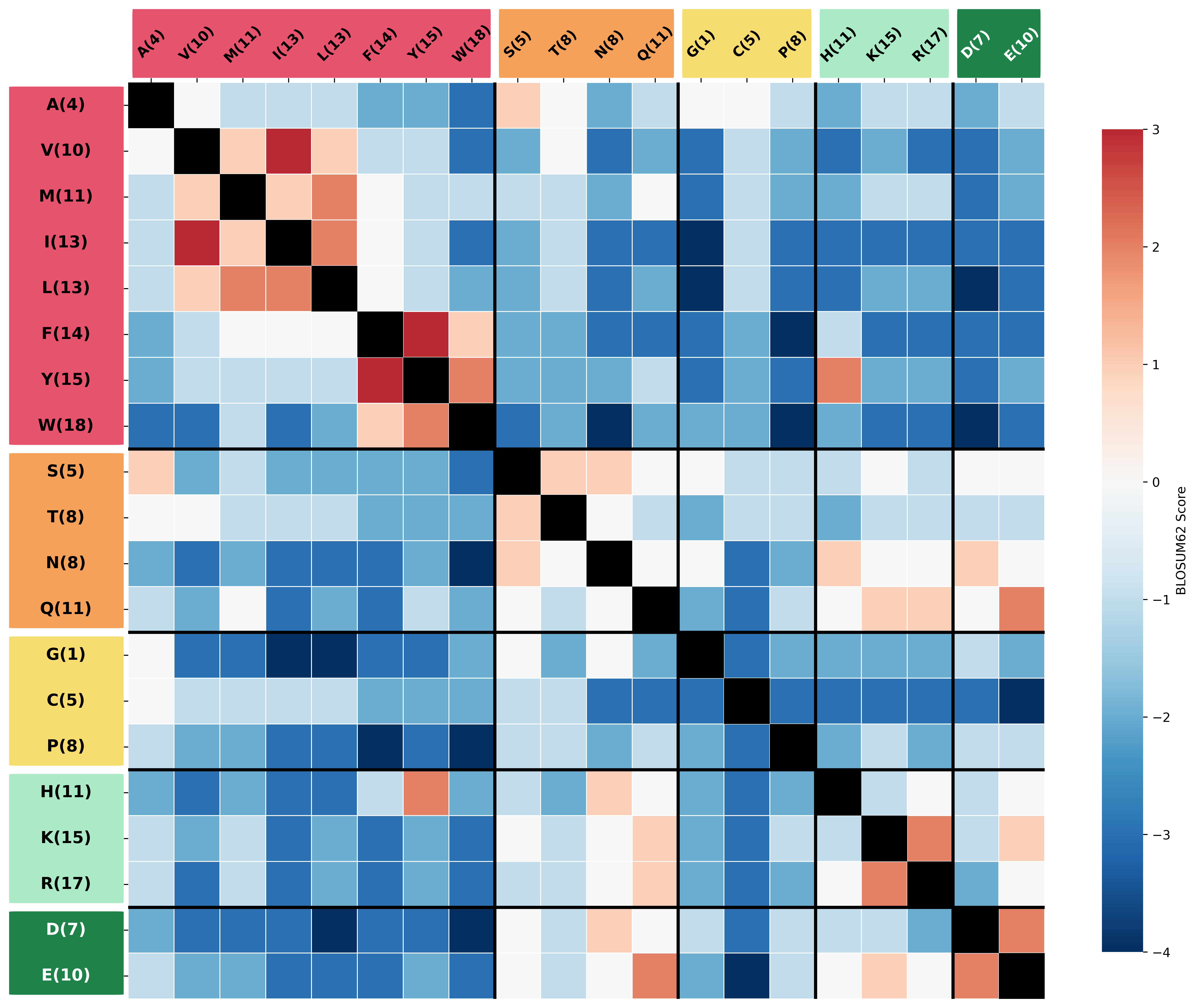
A few patterns jump out. Within each property group, most substitutions score near zero or positive, especially between amino acids of similar size: the branched-chain hydrophobic residues (Val, Ile, Leu) interchange readily, as do Asp and Glu (the two negatively charged residues) and Lys and Arg (two of the three positively charged ones). The hydrophobic aromatic residues (Phe, Tyr, Trp) also cluster together. Between groups, the scores turn negative: substituting a small hydrophobic residue for a charged one almost never happens in conserved positions, because the physicochemical mismatch is too large.
The most striking row in the matrix belongs to tryptophan. Trp is the largest and rarest amino acid, and its substitution scores are among the most negative in the matrix. When a tryptophan appears in a conserved position in a protein, it’s almost always doing something specific; evolution rarely allows it to be swapped out.
The Genetic Code
Substitution matrices tell us which amino acids are exchangeable in protein space, but there’s a second layer we haven’t touched: the DNA itself. Every substitution that makes it into a protein has to survive translation, which means it has to be reachable by a small number of point mutations in the underlying mRNA. The mapping from nucleotide triplets to amino acids, the genetic code, is what determines which substitutions are “close” in mutational space and which are far away.
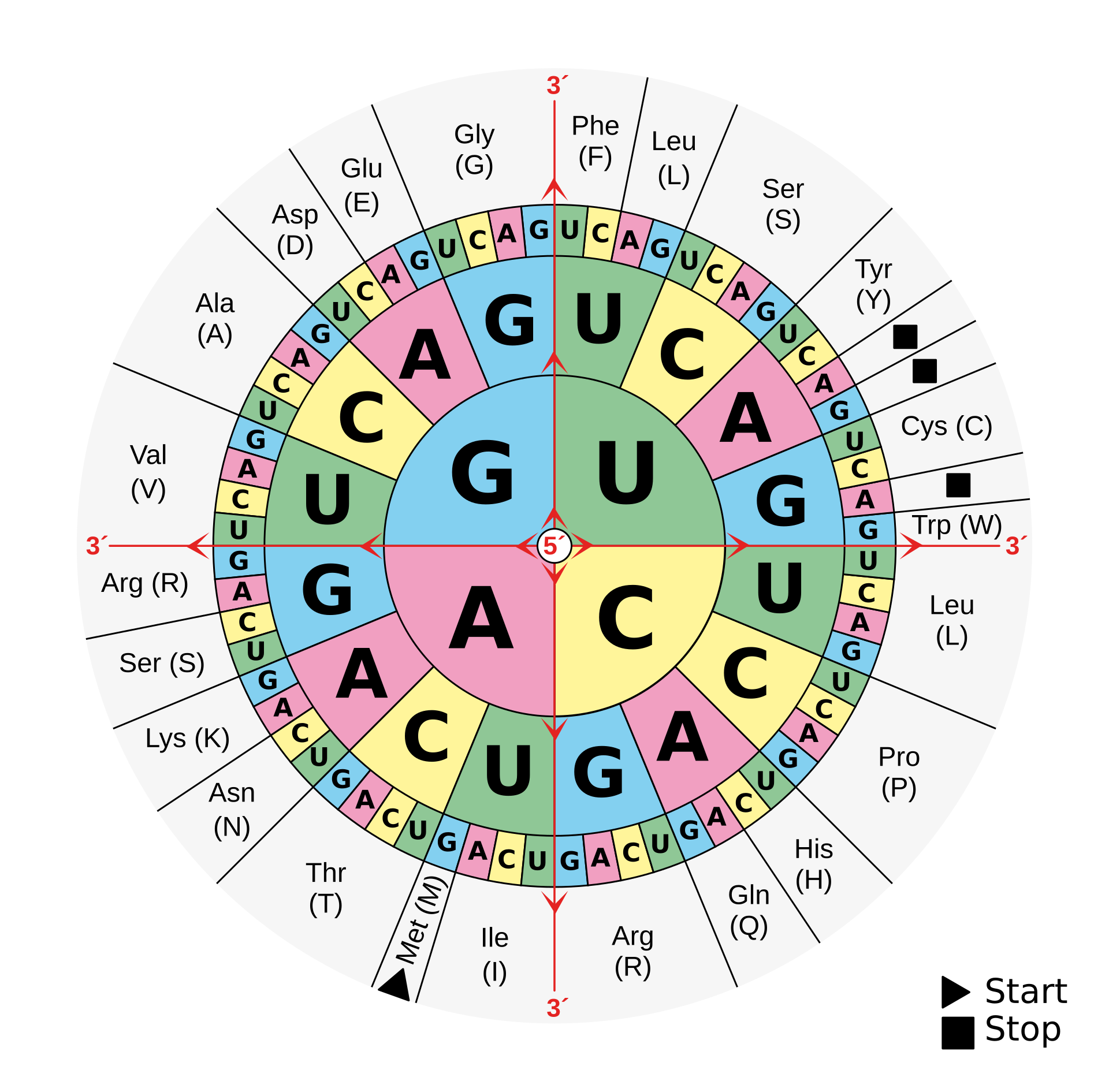
Three things stand out from the standard code. First, it is redundant: 61 sense codons map to 20 amino acids, so most amino acids are encoded by multiple codons (leucine, serine, and arginine each get six; only methionine and tryptophan get exactly one). Second, the redundancy is concentrated at the third position of the codon; changing the third base often leaves the amino acid unchanged. This is the wobble position, and it acts as a buffer against point mutations in DNA: a random mutation at the third position is the mutation most likely to be silent. Third, even when a point mutation does change the amino acid, the code is structured so that the replacement is usually chemically similar. Mutations at the first position tend to swap hydrophobic residues for other hydrophobic residues; mutations at the second position are the most likely to cause a radical change in physicochemical properties, but even then the code is biased toward minimizing the damage.
We can quantify this directly. For each pair of amino acids, we can count how many ways a single-nucleotide substitution in any of their codons converts one into the other. This gives us a 20×20 connectivity matrix that reflects which amino acids are “mutational neighbors.”
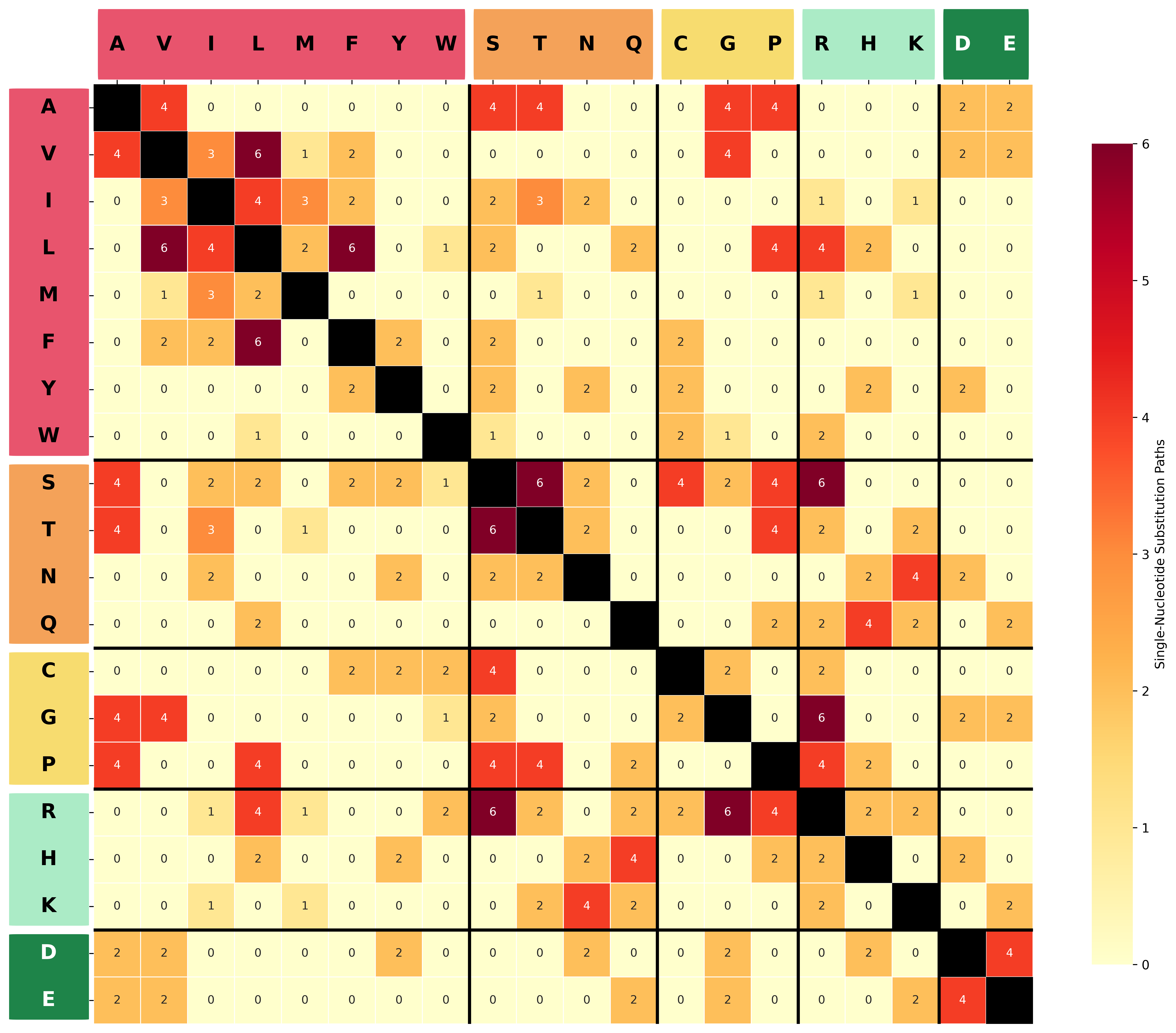
The connectivity matrix has a clear block-diagonal structure. Most single-nucleotide paths stay within a property group, which means the genetic code has been sculpted (by natural selection, or by the ancient history of which tRNAs matched which codons) to minimize the physicochemical impact of a random mutation. This “error-minimizing” property is sometimes called the genetic code’s robustness, and it is widely considered one of the strongest pieces of evidence that the code is not arbitrary; it has been optimized.
The alignment between substitution matrices and mutational proximity is not accidental. The substitution matrices we saw in the previous section are measured from real protein evolution, which is the combined outcome of (a) which mutations occur at the DNA level and (b) which mutations survive selection at the protein level. The genetic code’s error-minimizing layout means that (a) already pre-filters for physicochemical similarity, and selection imposes the remaining constraint.
* Why alanine and not glycine, the simplest amino acid? Glycine's lack of any side chain gives it unusual backbone flexibility, so substituting glycine would change the protein's conformational dynamics, not just remove the side chain's chemistry. Alanine's methyl group constrains the backbone like a normal amino acid while contributing almost nothing chemically.